Showing posts with label python. Show all posts
Showing posts with label python. Show all posts
Sunday, 8 June 2014
Running statistics using RunningStats (how to run on Python using SWIG and Distutils)
There is a more accurate method for calculating focal statistics than usually implemented in numerical packages. This is documented here and the C++ code is download from here
The SWIG setup file is simple:
which you run using:
The Distutils setup script is:
compile using "python setup.py install"
Then in python, a simple implementation of calculating the global stats from an array would be something like:
To show the convergence of locals means on the global mean, you might use the RunningStats class in the following way:
Finally, to compute running averages on the array you might use something like the following, where the function to window the data efficiently was downloaded from here
Obviously, as well as the mean, the above could be used for variance, standard deviation, kurtosis and skewness.
Tuesday, 15 October 2013
Sunday, 13 October 2013
How to pass arguments to python programs ...
... the right way (in my opinion). This is the basic template I always use. Not complicated, but the general formula might be useful to someone.
It allows you to pass arguments to a program with the use of flags, so they can be provided in any order. It has a 'help' flag (-h), will raise an error if no arguments are passed.
In this example one of the inputs (-a) is supposed to be a file. If the file is not provided, it uses Tkinter to open a simple 'get file' window dialog.
It then parses the other arguments out. Here I have assumed one argument is a string, one is a float, and one is an integer. If any arguments are not provided, it creates variables based on default variables. It prints everything it's doing to the screen.
It allows you to pass arguments to a program with the use of flags, so they can be provided in any order. It has a 'help' flag (-h), will raise an error if no arguments are passed.
In this example one of the inputs (-a) is supposed to be a file. If the file is not provided, it uses Tkinter to open a simple 'get file' window dialog.
It then parses the other arguments out. Here I have assumed one argument is a string, one is a float, and one is an integer. If any arguments are not provided, it creates variables based on default variables. It prints everything it's doing to the screen.
Saturday, 12 October 2013
Kivy python script for capturing displaying webcam
Kivy is very cool. It allows you to make graphical user interfaces for computers, tablets and smart phones ... in Python. Which is great for people like me. And you, I'm sure. And it's open source, of course.
Today was my first foray into programming using Kivy. Here was my first 'app'. It's not very sophisticated, but I find Kivy's documentation rather impenetrable and frustratingly lacking in examples. Therefore I hope more people like me post their code and projects on blogs.
Right, it detects and displays your webcam on the screen, draws a little button in the bottom. When you press that button it takes a screengrab and writes it to an image file. Voila! Maximise the window before you take a screen shot for best effects.
Today was my first foray into programming using Kivy. Here was my first 'app'. It's not very sophisticated, but I find Kivy's documentation rather impenetrable and frustratingly lacking in examples. Therefore I hope more people like me post their code and projects on blogs.
Right, it detects and displays your webcam on the screen, draws a little button in the bottom. When you press that button it takes a screengrab and writes it to an image file. Voila! Maximise the window before you take a screen shot for best effects.
Simple Python script to write a list of coordinates to a .geojson file
This has probably been done better elsewhere, but here's a really simple way to write a geojson file programmatically to display a set of coordinates, without having to bother learning the syntax of yet another python library. Geojson is a cool little format, open source and really easy.
Below, arrays x and y are decimal longitude and latitude respectively. The code would have to be modified to include a description for each coordinate, which could be included easily on line 15 if in the form of an iterable list.
Below, arrays x and y are decimal longitude and latitude respectively. The code would have to be modified to include a description for each coordinate, which could be included easily on line 15 if in the form of an iterable list.
Wednesday, 14 August 2013
Doing Spectral Analysis on Images with GMT
Mostly an excuse to play with, and show off the capabilities of, the rather fabulous Ipython notebook! I have created an Ipython notebook to demonstrate how to use GMT to do spectral analysis on images.
This is demonstrated by pulling wavelengths from images of rippled sandy sediments.
GMT is command-line mapping software for Unix/Linux systems. It's not really designed for spectral analysis in mind, especially not on images, but it is actually a really fast and efficient way of doing it!
This also requires the convert facility provided by Image Magick. Another bodacious tool I couldn't function without.
My notebook can be found here.
Fantastic images of ripples provided by http://skeptic.smugmug.com
This is demonstrated by pulling wavelengths from images of rippled sandy sediments.
GMT is command-line mapping software for Unix/Linux systems. It's not really designed for spectral analysis in mind, especially not on images, but it is actually a really fast and efficient way of doing it!
This also requires the convert facility provided by Image Magick. Another bodacious tool I couldn't function without.
My notebook can be found here.
Fantastic images of ripples provided by http://skeptic.smugmug.com
Labels:
bash,
data analysis,
GMT,
ipython,
photography,
python
Wednesday, 24 July 2013
Friday, 8 February 2013
Raspberry pi launch program on boot, and from desktop
The following is an example of how to launch a program running in a terminal upon boot, in Raspbian wheezy:
and add:
This will allow the same program to be launched from the desktop upon double-click:
add
and add:
This will allow the same program to be launched from the desktop upon double-click:
add
Saturday, 26 January 2013
Automatic Clustering of Geo-tagged Images. Part 2: Other Feature Metrics
In the last post I took a look at a simple method for trying to automatically cluster a set of images (of Horseshoe Bend, Glen Canyon, Arizona). Some of those picture were of the river (from different perspectives) and some of other things nearby.
With a goal of trying to separate the two classes, the results were reasonably satisfactory with image histograms as a feature metric, and a little user input. However, it'd be nice to get a more automated/objective way to separate the images into two discrete clusters. Perhaps the feature detection method requires a little more scrutiny?
In this post I look at 5 methods using the functionality of the excellent Mahotas toolbox for python downloaded here:
1) Image moments (I compute the first 5)
2) Haralick texture descriptors which are based on the co-occurrence matrix of the image
3) Zernike Moments
4) Threshold adjacency statistics (TAS)
5) Parameter-free threshold adjacency statistics (PFTAS)
The code is identical to the last post except for the feature extraction loop. What I'm looking for is as many images of the river at either low distances (extreme left in the dendrogram) or very high distances (extreme right in the dendrograms). These clusters have been highlighted below. There are 18 images of the river bend.
In order of success (low to high):
1) Zernike (using a radius of 50 pixels and 8 degrees, but other combinations tried):
This does a poor job, with no clusters at either end of the dendrogram.
2) Haralick:
There is 1 cluster of 4 images at the start
3) Moments:
There is 1 cluster of 6 images at the start, and 1 cluster of 2 at the end
4) TAS:
There is 1 cluster of 7 images at the start, and 1 cluster of 5 near the end, and both clusters are on the same stem
5) PFTAS:
The cluster at the end contains 16 out of 18 images of the river bend at the end. Plus there are no user-defined parameters. My kind of algorithm. The winner by far!
So there you have it, progress so far! No methods tested so far is perfect. There may or may not be a 'part 3' depending on my progress, or lack of!
With a goal of trying to separate the two classes, the results were reasonably satisfactory with image histograms as a feature metric, and a little user input. However, it'd be nice to get a more automated/objective way to separate the images into two discrete clusters. Perhaps the feature detection method requires a little more scrutiny?
In this post I look at 5 methods using the functionality of the excellent Mahotas toolbox for python downloaded here:
1) Image moments (I compute the first 5)
2) Haralick texture descriptors which are based on the co-occurrence matrix of the image
3) Zernike Moments
4) Threshold adjacency statistics (TAS)
5) Parameter-free threshold adjacency statistics (PFTAS)
The code is identical to the last post except for the feature extraction loop. What I'm looking for is as many images of the river at either low distances (extreme left in the dendrogram) or very high distances (extreme right in the dendrograms). These clusters have been highlighted below. There are 18 images of the river bend.
In order of success (low to high):
1) Zernike (using a radius of 50 pixels and 8 degrees, but other combinations tried):
2) Haralick:
There is 1 cluster of 4 images at the start
3) Moments:
4) TAS:
5) PFTAS:
The cluster at the end contains 16 out of 18 images of the river bend at the end. Plus there are no user-defined parameters. My kind of algorithm. The winner by far!
So there you have it, progress so far! No methods tested so far is perfect. There may or may not be a 'part 3' depending on my progress, or lack of!
Labels:
clustering,
data analysis,
image analysis,
python
Tuesday, 22 January 2013
Automatic Clustering of Geo-tagged Images. Part 1: using multi-dimensional histograms
There's a lot of geo-tagged images on the web. Sometimes the image coordinate is slightly wrong, or the scene isn't quite what you expect. It's therefore useful to have a way to automatically download images from a given place (specified by a coordinate) from the web, and automatically classify them according to content/scene.
In this post I describe a pythonic way to:
1) automatically download images based on input coordinate (lat, long)
2) extract a set features from each image
3) classify each image into groups
4) display the results as a dendrogram
This first example, 2) is achieved using a very simple means, namely the image histogram of image values. This doesn't take into account any texture similarities or connected components, etc. Nonetheless, it does a reasonably good job at classifying the images into a number of connected groups, as we shall see.
In subsequent posts I'm going to play with different ways to cluster images based on similarity, so watch this space!
User inputs: give a lat and long of a location, a path where you want the geo-tagged photos, and the number of images to download
First, import the libraries you'll need. Then interrogates the website and downloads the images.
As you can see the resulting images are a mixed bag. There's images of the river bend, the road, the desert, Lake Powell and other random stuff. My task is to automatically classify these images so it's easier to pull out just the images of the river bend
The following compiles a list of these images. The last part is to sort which is not necessary but doing so converts the list into a numpy array which is. Then clustering of the images is achieved using Jan Erik Solem's rather wonderful book 'Programming Computer Vision with Python'. The examples from which can be downloaded here. Download this one, then this bit of code does the clustering:
In this post I describe a pythonic way to:
1) automatically download images based on input coordinate (lat, long)
2) extract a set features from each image
3) classify each image into groups
4) display the results as a dendrogram
This first example, 2) is achieved using a very simple means, namely the image histogram of image values. This doesn't take into account any texture similarities or connected components, etc. Nonetheless, it does a reasonably good job at classifying the images into a number of connected groups, as we shall see.
In subsequent posts I'm going to play with different ways to cluster images based on similarity, so watch this space!
User inputs: give a lat and long of a location, a path where you want the geo-tagged photos, and the number of images to download
First, import the libraries you'll need. Then interrogates the website and downloads the images.
As you can see the resulting images are a mixed bag. There's images of the river bend, the road, the desert, Lake Powell and other random stuff. My task is to automatically classify these images so it's easier to pull out just the images of the river bend
The following compiles a list of these images. The last part is to sort which is not necessary but doing so converts the list into a numpy array which is. Then clustering of the images is achieved using Jan Erik Solem's rather wonderful book 'Programming Computer Vision with Python'. The examples from which can be downloaded here. Download this one, then this bit of code does the clustering:
The approach taken is to use hierarchical clustering using a simple euclidean distance function. This bit of code does the dendogram plot of the images:
which in my case looks like this (click on the image to see the full extent):
It's a little small, but you can just about see (if you scroll to the right) that it does a good job at clustering images of the river bend which look similar. The single-clustered, or really un-clustered, images to the left are those of the rim, road, side walls, etc which don't look anything like the river bend.
Next, reorder the image list in terms of similarity distance (which increases in both the x and y directions of the dendrogram above)
Which gives me:
As you can see they are all images of the river bend, except the 2nd from the left on the top row, which is a picture of a shrub. Interestingly, the pattern of a circular shrub surrounded by a strip of sand is visually similar to the horse shoe bend!!
However, we don't want to include it with images of the river, which is why a more sophisticated method that image histogram is required to classify and group similar image ... the subject of a later post.
Labels:
clustering,
data analysis,
image analysis,
python
Sunday, 20 January 2013
Alpha Shapes in Python
Alpha shapes include convex and concave hulls. Convex hull algorithms are ten a penny, so what we're really interested in here in the concave hull of an irregularly or otherwise non-convex shaped 2d point cloud, which by all accounts is more difficult.
The function is here:
The above is essentially the same wrapper as posted here, except a different way of reading in the data, and providing the option to specify a probe radius. It uses Ken Clarkson's C-code, instructions on how to compile are here
An example implementation:
The above data generation is translated from the example in a matlab alpha shape function . Then:
Which produces (points are blue dots, alpha shape is red line):
and on a larger data set (with radius=1, this is essentially the convex hull):
and a true concave hull:
The function is here:
The above is essentially the same wrapper as posted here, except a different way of reading in the data, and providing the option to specify a probe radius. It uses Ken Clarkson's C-code, instructions on how to compile are here
An example implementation:
The above data generation is translated from the example in a matlab alpha shape function . Then:
Which produces (points are blue dots, alpha shape is red line):
and on a larger data set (with radius=1, this is essentially the convex hull):
and a true concave hull:
Labels:
alpha shapes,
data analysis,
hull,
python,
voronoi tessellation
Patchwork quilt in Python
Here's a fun python function to make a plot of [x,y] coordinate data as a patchwork quilt with user-defined 'complexity'. It initially segments the data into 'numsegs' number of clusters by a k-means algorithm. It then takes each segment and creates 'numclass' sub-clusters based on the euclidean distance from the centroid of the cluster. Finally, it plots each subcluster a different colour and prints the result as a png file.
inputs:
'data' is a Nx2 numpy array of [x,y] points
'numsegs' and 'numclass' are integer scalars. The greater these numbers the greater the 'complexity' of the output and the longer the processing time
Some example outputs in increasing complexity:
numsegs=10, numclass=5
numsegs=15, numclass=15
numsegs=20, numclass=50
inputs:
'data' is a Nx2 numpy array of [x,y] points
'numsegs' and 'numclass' are integer scalars. The greater these numbers the greater the 'complexity' of the output and the longer the processing time
Some example outputs in increasing complexity:
numsegs=10, numclass=5
numsegs=15, numclass=15
numsegs=20, numclass=50
Sunday, 9 September 2012
Craigslist Part 2: using matlab to plot your search results in Google Maps
In 'Craigslist Part 1' I demonstrated a way to use matlab to automatically search craigslist for properties with certain attributes. In this post I show how to use matlab and python to create a kml file for plotting the results in Google Earth.
Download the matlab googleearth toolbox from here
Add google earth toolbox to path. Import search results data and get map links (location strings). Loop through each map location and string the address from the url, then geocode to obtain coordinates.
The function 'geocode' writes and executes a python script to geocode the addresses (turn the address strings into longitude and latitude coordinates). The python module may be downloaded here
Once we have the coordinates, we then need to get rid of nans and outliers (badly converted coordinates due to unreadable address strings). Use the google earth toolbox to build the kml file. Finally, run google earth and open the kml file using a system command:
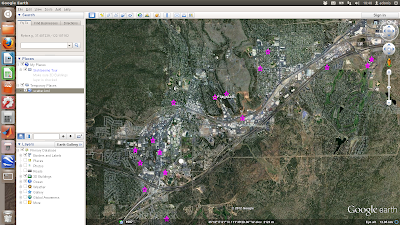 The above is a simple scatter plot which only shows the location of the properties and not any information about them. Next shows a more complicated example where the points are plotted with labels (the asking price) and text details (the google map links) in pop-up boxes
First each coordinate pair is packaged with the map and name tags. Concatenate the strings for each coordinate and make a kml file. Finally, run google earth and open the kml file using a system command:
The above is a simple scatter plot which only shows the location of the properties and not any information about them. Next shows a more complicated example where the points are plotted with labels (the asking price) and text details (the google map links) in pop-up boxes
First each coordinate pair is packaged with the map and name tags. Concatenate the strings for each coordinate and make a kml file. Finally, run google earth and open the kml file using a system command:
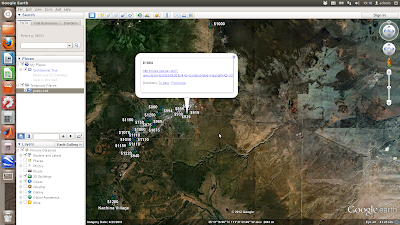
 The above is a simple scatter plot which only shows the location of the properties and not any information about them. Next shows a more complicated example where the points are plotted with labels (the asking price) and text details (the google map links) in pop-up boxes
First each coordinate pair is packaged with the map and name tags. Concatenate the strings for each coordinate and make a kml file. Finally, run google earth and open the kml file using a system command:
The above is a simple scatter plot which only shows the location of the properties and not any information about them. Next shows a more complicated example where the points are plotted with labels (the asking price) and text details (the google map links) in pop-up boxes
First each coordinate pair is packaged with the map and name tags. Concatenate the strings for each coordinate and make a kml file. Finally, run google earth and open the kml file using a system command:

Labels:
craigslist,
google earth,
internet,
matlab,
python
Saturday, 10 September 2011
Ellipse Fitting to 2D points, part 2: Python
The second of 3 posts presenting algorithms for the ellipsoid method of Khachiyan for 2D point clouds. This version in python will return: 1) the ellipse radius in x, 2) the ellipse radius in y
x is a 2xN array [x,y] points
Saturday, 26 March 2011
Controlling a serial device with Python
This program to sends a command to serial device, receives data from the device, converts it to numeric format, and graphs the data in real time. It is written in Python using wxPython for the display.
The device is connected to /dev/ttyS0 with a baud rate of 19200, 8-bit data, no parity, and 1 stopbit. You need to edit the line 'serial.Serial' with the relevant parameters to use this code for your device. Editing 'DATA_LENGTH' will change the rate at which the graph scrolls forward. Also, you'd need to change the 'command' that the device understands as 'send me data' (here it is the s character - line 'input'). Knowledge of the format of the returned data is also essential (here I turn ascii into 2 integers) line 'struct.unpack' - see here for help). Otherwise this should work with any device. Tested on Ubuntu 9.10 and 10.4 using serial port and USB-serial converters.
The device is connected to /dev/ttyS0 with a baud rate of 19200, 8-bit data, no parity, and 1 stopbit. You need to edit the line 'serial.Serial' with the relevant parameters to use this code for your device. Editing 'DATA_LENGTH' will change the rate at which the graph scrolls forward. Also, you'd need to change the 'command' that the device understands as 'send me data' (here it is the s character - line 'input'). Knowledge of the format of the returned data is also essential (here I turn ascii into 2 integers) line 'struct.unpack' - see here for help). Otherwise this should work with any device. Tested on Ubuntu 9.10 and 10.4 using serial port and USB-serial converters.
Thursday, 16 April 2009
Frequency Transforms on Images
As part of my work, I have to do a lot of spectral analysis with images. Traditionally I have used MATLAB to do this - it is exceptionally easy and powerful, and works just fine. As a simple example, if I want a power-spectrum of an image, treated as a 2D matrix of 8-bit numbers, the sequence of commands is this:
Seven lines of commands. Couldn't be simpler. That's the beauty of MATLAB. The snag is that it costs several hundreds to thousands of dollars for the program, which makes it difficult to share your code with other people. So recently I've thought about coding up some things I want to share with other people in Python.
Python is a cross-platform, open-source programming language and environment. It is not too dissimilar to MATLAB in many respects, especially when you install the right libraries. In order to do the same in Python as above, as easy as possible, I needed to install Python Imaging Library (PIL), NumPy and SciPy. Then the sequence of operations is as follows:
Same answer. Not bad - not harder, just different. And completely free. Oh, and this'll work on UNIX, Linux, Mac, and PC when Python is installed (with a couple of extra libraries). Great!
%------------------- MATLAB
% read image in, convert to greyscale and double-precision
image=double(rgb2gray(imread('myimage.jpg')));
% crop to smallest dimension
[m,n]=size(im);
im=im(1:min(m,n),1:min(m,n));
% find image mean
imMean=mean(im(:));
% and subtract mean from the image
demean=im-imMean;
% find the 2D fourier transform
ft=fft2(demean);
% power spectrum is the element-by-element square of the fourier transform
spectrum=abs(ft).^2;
Seven lines of commands. Couldn't be simpler. That's the beauty of MATLAB. The snag is that it costs several hundreds to thousands of dollars for the program, which makes it difficult to share your code with other people. So recently I've thought about coding up some things I want to share with other people in Python.
Python is a cross-platform, open-source programming language and environment. It is not too dissimilar to MATLAB in many respects, especially when you install the right libraries. In order to do the same in Python as above, as easy as possible, I needed to install Python Imaging Library (PIL), NumPy and SciPy. Then the sequence of operations is as follows:
# ----------------PYTHON
#-----------------------
# declare libraries to use
import Image
import scipy.fftpack
from pylab import*
import cmath
from numpy import*
# load image, convert to greyscale
im=Image.open('myimage.jpg').convert("L")
# crop to smallest dimension
box=(0,0,im.ny,im.nx)
region=im.crop(box)
# find mean
imMean=mean(asarray(region))
# 2D fourier transform of demeaned image
ft=fft2(region-imMean)
# power spectrum
spectrum=pow(abs(ft),2)
Same answer. Not bad - not harder, just different. And completely free. Oh, and this'll work on UNIX, Linux, Mac, and PC when Python is installed (with a couple of extra libraries). Great!
Hmm, I thought, I wonder if it is possible to do the same in a piece of non-numerical software. I tried ImageMagick because its free and ubiquitous on Linux and Mac, and also available for PC, and it has a massive user-community. It is incredibly powerful as a command-line graphics package - even more powerful than PhotoShop and Illustrator, and like all UNIX-based applications, infinitely flexible. It turns out it is a little more tricky to calculate the power spectrum in ImageMagick, but here's how.
First, you need the right tools - ImageMagick is usually already present on most Linux distributions - if not, if a debian/Ubuntu user use sudo apt-get install imagemagick, or download from the webiste hyperlinked above. Then you'll need an open-source C-program for calculation of FFTs: this is provided by FTW - download fftw-3.2.1.tar.gz, untar, cd to that directory, then install the binary by issuing the usual commands:
This C-program is no good on its own, so the last ingredient is a wrapper for ImageMagick. Fortunately - nay, crucially, this has been provided by Sean Burke and Fred Weinhaus - introducing IMFFT! The program can be downloaded and installed from here. And here is an in-depth tutorial of how to use it (I'm only just getting to grips with it myself). For some reason the compiled program is called 'demo'. I move it to my /bin directory so my computer can see it wherever I am working, and in the process change the name to something more suitable - fft:
Right, it's a little bit more involved, but here is the bash shell script for the power spectrum in ImageMagick:
Voila! The spectrum is the image 'ggc_dm_F_Q16_spec.tiff'. I haven't yet worked out how to keep all the data inside the program, to eliminate the need (and cumulative errors associated with) writing the outputs at every stage to file, and eading them back in again. This means I can't use the spectrum for any scientific application. But in theory it should be possible. Any suggestions welcome!
./configure
sudo make && make install
This C-program is no good on its own, so the last ingredient is a wrapper for ImageMagick. Fortunately - nay, crucially, this has been provided by Sean Burke and Fred Weinhaus - introducing IMFFT! The program can be downloaded and installed from here. And here is an in-depth tutorial of how to use it (I'm only just getting to grips with it myself). For some reason the compiled program is called 'demo'. I move it to my /bin directory so my computer can see it wherever I am working, and in the process change the name to something more suitable - fft:
cp demo ./fft && sudo mv ./fft /bin
Right, it's a little bit more involved, but here is the bash shell script for the power spectrum in ImageMagick:
#----------------------------- IMAGEMAGICK
#-------------------------------------------------
#!/bin/bash
# read image and convert to greyscale
convert myimage.tif -colorspace gray input_grey.tiff
# find min dimension for cropping
min=`convert input_grey.tiff -format "%[fx: min(w,h)]" info:`
# crop image
convert input_grey.tiff -background white -gravity center -extent ${min}x${min} input_grey_crop.tiff
# de-mean image
# get data
data=`convert input_grey_crop.tiff -verbose info:`
# find mean as proportion
mean=`echo "$data" | sed -n '/^.*[Mm]ean:.*[(]\([0-9.]*\).*$/{ s//\1/; p; q; }'`
# convert mean to percent
m=`echo "scale=1; $mean *100 / 1" | bc`
# de-mean image
convert input_grey_crop.tiff -evaluate subtract ${m}% ggc_dm.tiff
# pad to power of 2
p2=`convert ggc_dm.tiff -format "%[fx:2^(ceil(log(max(w,h))/log(2)))]" info:`
convert ggc_dm.tiff -background white -gravity center -extent ${p2}x${p2} ggc_dm_pad.tiff
# forward fast fourier transform
fft f ggc_dm_pad.tiff
# creates ggc_dm_F_Q16.tiff, on which get spectrum
convert ggc_dm_F_Q16.tiff -contrast-stretch 0% -gamma .5 -fill "rgb(1,1,1)" -opaque black ggc_dm_F_Q16_spec.tiff
Voila! The spectrum is the image 'ggc_dm_F_Q16_spec.tiff'. I haven't yet worked out how to keep all the data inside the program, to eliminate the need (and cumulative errors associated with) writing the outputs at every stage to file, and eading them back in again. This means I can't use the spectrum for any scientific application. But in theory it should be possible. Any suggestions welcome!
Subscribe to:
Posts (Atom)















